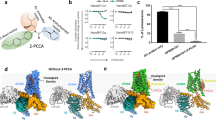
Abstract
At least 120 non-olfactory G-protein-coupled receptors in the human genome are ‘orphans’ for which endogenous ligands are unknown, and many have no selective ligands, hindering the determination of their biological functions and clinical relevance. Among these is GPR68, a proton receptor that lacks small molecule modulators for probing its biology. Using yeast-based screens against GPR68, here we identify the benzodiazepine drug lorazepam as a non-selective GPR68 positive allosteric modulator. More than 3,000 GPR68 homology models were refined to recognize lorazepam in a putative allosteric site. Docking 3.1 million molecules predicted new GPR68 modulators, many of which were confirmed in functional assays. One potent GPR68 modulator, ogerin, suppressed recall in fear conditioning in wild-type but not in GPR68-knockout mice. The same approach led to the discovery of allosteric agonists and negative allosteric modulators for GPR65. Combining physical and structure-based screening may be broadly useful for ligand discovery for understudied and orphan GPCRs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Roth, B. L. & Kroeze, W. K. Integrated approaches for genome-wide interrogation of the druggable non-olfactory G protein-coupled receptor superfamily. J. Biol. Chem. 290, 19471–19477 (2015)
Davenport, A. P. et al. International Union of Basic and Clinical Pharmacology. LXXXVIII. G protein-coupled receptor list: recommendations for new pairings with cognate ligands. Pharmacol. Rev. 65, 967–986 (2013)
Chung, S., Funakoshi, T. & Civelli, O. Orphan GPCR research. Br. J. Pharmacol . 153 (suppl. 1), S339–S346 (2008)
Knapp, S. et al. A public–private partnership to unlock the untargeted kinome. Nature Chem. Biol. 9, 3–6 (2013)
Ferguson, F. M. et al. Targeting low-druggability bromodomains: fragment based screening and inhibitor design against the BAZ2B bromodomain. J. Med. Chem. 56, 10183–10187 (2013)
Leung, D., Hardouin, C., Boger, D. L. & Cravatt, B. F. Discovering potent and selective reversible inhibitors of enzymes in complex proteomes. Nature Biotechnol. 21, 687–691 (2003)
Ludwig, M. G. et al. Proton-sensing G-protein-coupled receptors. Nature 425, 93–98 (2003)
Mogi, C. et al. Sphingosylphosphorylcholine antagonizes proton-sensing ovarian cancer G-protein-coupled receptor 1 (OGR1)-mediated inositol phosphate production and cAMP accumulation. J. Pharmacol. Sci. 99, 160–167 (2005)
Li, J. et al. Ovarian cancer G protein coupled receptor 1 suppresses cell migration of MCF7 breast cancer cells via a Gα12/13-Rho-Rac1 pathway. J. Mol. Signal. 8, 6 (2013)
Singh, L. S. et al. Ovarian cancer G protein-coupled receptor 1, a new metastasis suppressor gene in prostate cancer. J. Natl. Cancer Inst. 99, 1313–1327 (2007)
Schneider, J. W. et al. Coupling hippocampal neurogenesis to brain pH through proneurogenic small molecules that regulate proton sensing G protein-coupled receptors. ACS Chem. Neurosci . 3, 557–568 (2012)
Frick, K. K., Krieger, N. S., Nehrke, K. & Bushinsky, D. A. Metabolic acidosis increases intracellular calcium in bone cells through activation of the proton receptor OGR1. J. Bone Miner. Res. 24, 305–313 (2009)
Komarova, S. V., Pereverzev, A., Shum, J. W., Sims, S. M. & Dixon, S. J. Convergent signaling by acidosis and receptor activator of NF-κB ligand (RANKL) on the calcium/calcineurin/NFAT pathway in osteoclasts. Proc. Natl Acad. Sci. USA 102, 2643–2648 (2005)
Yang, M. et al. Expression of and role for ovarian cancer G-protein-coupled receptor 1 (OGR1) during osteoclastogenesis. J. Biol. Chem. 281, 23598–23605 (2006)
Russell, J. L. et al. Regulated expression of pH sensing G protein-coupled receptor-68 identified through chemical biology defines a new drug target for ischemic heart disease. ACS Chem. Biol. 7, 1077–1083 (2012)
Mohebbi, N. et al. The proton-activated G protein coupled receptor OGR1 acutely regulates the activity of epithelial proton transport proteins. Cell. Physiol. Biochem. 29, 313–324 (2012)
Regard, J. B., Sato, I. T. & Coughlin, S. R. Anatomical profiling of G protein-coupled receptor expression. Cell 135, 561–571 (2008)
Chen, Y. J., Huang, C. W., Lin, C. S., Chang, W. H. & Sun, W. H. Expression and function of proton-sensing G-protein-coupled receptors in inflammatory pain. Mol. Pain 5, 39 (2009)
Saxena, H. et al. The GPCR OGR1 (GPR68) mediates diverse signalling and contraction of airway smooth muscle in response to small reductions in extracellular pH. Br. J. Pharmacol. 166, 981–990 (2012)
Wang, J., Sun, Y., Tomura, H. & Okajima, F. Ovarian cancer G-protein-coupled receptor 1 induces the expression of the pain mediator prostaglandin E2 in response to an acidic extracellular environment in human osteoblast-like cells. Int. J. Biochem. Cell Biol. 44, 1937–1941 (2012)
Li, H. et al. Abnormalities in osteoclastogenesis and decreased tumorigenesis in mice deficient for ovarian cancer G protein-coupled receptor 1. PLoS ONE 4, e5705 (2009)
Aoki, H. et al. Proton-sensing ovarian cancer g protein-coupled receptor 1 on dendritic cells is required for airway responses in a murine asthma model. PLoS ONE 8, e79985 (2013)
Mogi, C., Nakakura, T. & Okajima, F. Role of extracellular proton-sensing OGR1 in regulation of insulin secretion and pancreatic β-cell functions. Endocr. J. 61, 101–110 (2013)
Okajima, F. Regulation of inflammation by extracellular acidification and proton-sensing GPCRs. Cell. Signal. 25, 2263–2271 (2013)
Dong, S., Rogan, S. C. & Roth, B. L. Directed molecular evolution of DREADDs: a generic approach to creating next-generation RASSLs. Nature Protocols 5, 561–573 (2010)
Mysinger, M. M. & Shoichet, B. K. Rapid context-dependent ligand desolvation in molecular docking. J. Chem. Inf. Model. 50, 1561–1573 (2010)
Evers, A. & Klebe, G. Ligand-supported homology modeling of G-protein-coupled receptor sites: models sufficient for successful virtual screening. Angew. Chem. Int. Ed. Engl. 43, 248–251 (2004)
Cavasotto, C. N. et al. Discovery of novel chemotypes to a G-protein-coupled receptor through ligand-steered homology modeling and structure-based virtual screening. J. Med. Chem. 51, 581–588 (2008)
Katritch, V., Rueda, M., Lam, P. C.-H., Yeager, M. & Abagyan, R. GPCR 3D homology models for ligand screening: lessons learned from blind predictions of adenosine A2a receptor complex. Proteins 78, 197–211 (2010)
Leach, K., Sexton, P. M. & Christopoulos, A. Allosteric GPCR modulators: taking advantage of permissive receptor pharmacology. Trends Pharmacol. Sci. 28, 382–389 (2007)
Keiser, M. J. et al. Relating protein pharmacology by ligand chemistry. Nature Biotechnol. 25, 197–206 (2007)
Kalk, P. et al. The adenosine A1 receptor antagonist SLV320 reduces myocardial fibrosis in rats with 5/6 nephrectomy without affecting blood pressure. Br. J. Pharmacol. 151, 1025–1032 (2007)
Thompson, S.-A., Wingrove, P. B., Connelly, L., Whiting, P. J. & Wafford, K. A. Tracazolate reveals a novel type of allosteric interaction with recombinant γ-aminobutyric acidA receptors. Mol. Pharmacol. 61, 861–869 (2002)
Tomura, H. et al. Prostaglandin I2 production and cAMP accumulation in response to acidic extracellular pH through OGR1 in human aortic smooth muscle cells. J. Biol. Chem. 280, 34458–34464 (2005)
Ichimonji, I. et al. Extracellular acidification stimulates IL-6 production and Ca2+ mobilization through proton-sensing OGR1 receptors in human airway smooth muscle cells. Am. J. Physiol. Lung Cell. Mol. Physiol. 299, L567–L577 (2010)
Liu, J. P. et al. Ovarian cancer G protein-coupled receptor 1-dependent and -independent vascular actions to acidic pH in human aortic smooth muscle cells. Am. J. Physiol. Heart Circ. Physiol. 299, H731–H742 (2010)
Matsuzaki, S. et al. Extracellular acidification induces connective tissue growth factor production through proton-sensing receptor OGR1 in human airway smooth muscle cells. Biochem. Biophys. Res. Commun. 413, 499–503 (2011)
Gravius, A., Barberi, C., Schäfer, D., Schmidt, W. J. & Danysz, W. The role of group I metabotropic glutamate receptors in acquisition and expression of contextual and auditory fear conditioning in rats – a comparison. Neuropharmacology 51, 1146–1155 (2006)
Daumas, S. et al. Transient activation of the CA3 Kappa opioid system in the dorsal hippocampus modulates complex memory processing in mice. Neurobiol. Learn. Mem. 88, 94–103 (2007)
Phillips, R. G. & LeDoux, J. E. Differential contribution of amygdala and hippocampus to cued and contextual fear conditioning. Behav. Neurosci. 106, 274–285 (1992)
Onozawa, Y. et al. Activation of T cell death-associated gene 8 regulates the cytokine production of T cells and macrophages in vitro . Eur. J. Pharmacol. 683, 325–331 (2012)
Pompéia, S., Manzano, G. M., Tufik, S. & Bueno, O. F. What makes lorazepam different from other benzodiazepines? J. Physiol. (Lond.) 569, 709 (2005)
Greenblatt, D. J. et al. Clinical pharmacokinetics of lorazepam. I. Absorption and disposition of oral 14C-lorazepam. Clin. Pharmacol. Ther. 20, 329–341 (1976)
Schoepp, D. D. Where will new neuroscience therapies come from? Nature Rev . Drug Discov . 10, 715–716 (2011)
Eswar, N. et al. Comparative protein structure modeling using MODELLER. Curr. Protoc. Protein Sci . Chapter 2, Unit 2.9 (2001)
Yang, Q. & Sharp, K. A. Building alternate protein structures using the elastic network model. Proteins 74, 682–700 (2009)
Jacobson, M. P., Friesner, R. A., Xiang, Z. & Honig, B. On the role of the crystal environment in determining protein side-chain conformations. J. Mol. Biol. 320, 597–608 (2002)
Li, J., Zhu, T., Cramer, C. J. & Truhlar, D. G. New class IV charge model for extracting accurate partial charges from wave functions. J. Phys. Chem. A 102, 1820–1831 (1998)
Chambers, C. C., Hawkins, G. D., Cramer, C. J. & Truhlar, D. G. Model for aqueous solvation based on class IV atomic charges and first solvation shell effects. J. Phys. Chem. 100, 16385–16398 (1996)
Hert, J., Keiser, M. J., Irwin, J. J., Oprea, T. I. & Shoichet, B. K. Quantifying the relationships among drug classes. J. Chem. Inf. Model. 48, 755–765 (2008)
Gaulton, A. et al. ChEMBL: a large-scale bioactivity database for drug discovery. Nucleic Acids Res. 40, D1100–D1107 (2012)
Mumberg, D., Muller, R. & Funk, M. Yeast vectors for the controlled expression of heterologous proteins in different genetic backgrounds. Gene 156, 119–122 (1995)
Erlenbach, I. et al. Functional expression of M1, M3 and M5 muscarinic acetylcholine receptors in yeast. J. Neurochem. 77, 1327–1337 (2001)
Armbruster, B. N., Li, X., Pausch, M. H., Herlitze, S. & Roth, B. L. Evolving the lock to fit the key to create a family of G protein-coupled receptors potently activated by an inert ligand. Proc. Natl Acad. Sci. USA 104, 5163–5168 (2007)
Christopoulos, A. & Kenakin, T. G protein-coupled receptor allosterism and complexing. Pharmacol. Rev. 54, 323–374 (2002)
Besnard, J. et al. Automated design of ligands to polypharmacological profiles. Nature 492, 215–220 (2012)
Keiser, M. J. et al. Predicting new molecular targets for known drugs. Nature 462, 175–181 (2009)
Horvat, S. J. et al. A-kinase anchoring proteins regulate compartmentalized cAMP signaling in airway smooth muscle. FASEB J . 26, 3670–3679 (2012)
Huang, X.-P., Mangano, T., Hufeisen, S., Setola, V. & Roth, B. L. Identification of human ether-à-go-go related gene modulators by three screening platforms in an academic drug-discovery setting. Assay Drug Dev. Technol. 8, 727–742 (2010)
Acknowledgements
This work was supported by National Institutes of Health (NIH) grants U01104974 (B.L.R., B.K.S. and W.K.K.), R01 DA017204 (B.L.R. and W.K.K.) and the National Institute of Mental Health Psychoactive Drug Screening Program (NIMH PDSP) (X.-P.H., H.Z., M.S.F., W.K.K., T.J.M., A.J. and B.L.R.), the Michael Hooker Chair for Protein Therapeutics and Translational Proteomics to B.L.R.; Genentech Foundation Predoctoral Fellowship (J.K.); NIH grants GM59957 and GM71896 (B.K.S.) and the Structural Genomics Consortium (B.K.S.); grant P01 HL114471 (R.B.P. and D.A.D.); NICHD grant U54 HD079124 (M.S.M., K.A.S., V.N.); NIH grant U19MH082441 (B.L.R., J.J. and X.C.). We thank Mark Pausch (Merck & Co.) for providing us Gs- and Gq-yeast strains for yeast screening assays.
Author information
Authors and Affiliations
Contributions
X.-P.H. subcloned GPR68 for yeast screening, made GPR68 and GPR65 mutants, designed, carried out cell-based screening assays, analysed results, and wrote the paper. J.K. designed and developed homology models, carried out docking screens, analysed results, and wrote the paper. W.K.K. set up and performed yeast screening assays, analysed results, and wrote the paper. H.Z. and M.S.F. designed, performed in vivo fear-conditioning studies, analysed results, and wrote the paper. M.S.F. and B.L.R. dubbed ZINC67740571 ‘ogerin’. B.H.K. created the GPR68-knockout mice. S.S.M., K.A.S. and V.N. carried out initial phenotypic characterization, analysed results, and wrote the paper. X.C. and J.J. synthesized ZINC32547799, ZINC67740571 (ogerin) and ogerin analogues (compounds 33548–33561, C3 and C4) for functional assays and in vivo studies, and wrote the paper. T.J.M. carried out radioligand binding assays. A.J. prepared drug plates and plasmids for initial screening. R.B.P. and D.A.D. designed and carried out anti-haemagglutinin immunoblot assays, analysed results, and wrote the paper. S.W. designed primers, prepared Flag-tagged GPR68 wild-type and mutant plasmids, performed anti-Flag western blot assays, and analysed results. T.K. analysed results and wrote the paper. B.L.R. and B.K.S. coordinated and supervised the project, and with the other authors wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Validation and confirmation of GPCR activation assays.
a–o, Yeast (a–k) and HEK293T cell (l–o) GPCR activation assays. a–d, Concentration-dependent growth of GPR43-expressing Gs yeast (a), GPR43-expressing Gq yeast (b), GPR41-expressing Gs yeast (c), and GPR41-expressing Gq yeast (d) in response to various short-chain fatty acids (SCFAs). e–h, Concentration-dependent growth of GPR39-expressing Gs yeast (GPR39s) in response to zinc ions (e), chromium ions (f), cadmium ions (g) and iron ions (h). i–k, Concentration-dependent cAMP responses of GPR39-expressing HEK293T cells to ZnCl2 (i), ZnSO4 (j) or CdSO4 (k) as measured by luciferase cAMP reporter assay. l, N-unsubstituted benzodiazepines (lorazepam, clonazepam, desmethyldiazepam and norfludiazepam; 10 μM) stimulated cAMP production in a GPR68- and pH-dependent manner. Data are mean ± s.e.m. (n = 3–66 measurements). m–p, Concentration–response curves of N-unsubstituted benzodiazepines lorazepam (m), desmethyldiazepam (n), clonazepam (o) and norfludiazepam (p) at pH 6.50 or 7.40 in GPR68-transfected HEK293T cells (structures in Supplementary Table 1). Normalized results represent mean ± s.e.m. (n = 3) and curves were analysed in GraphPad Prism using the built-in 4 parameter logistic function.
Extended Data Figure 2 Lorazepam and ogerin have minimal GPR4 or GPR65 activity.
a–d, Effect of lorazepam (a, b) or ogerin (c, d) on GPR4 (a, c) or GPR65 (b, d); data represent normalized mean ± s.e.m. (n = 3). Sequencing alignment proton-sensing receptor and docking poses for ogerin and its analogues. e, GPR68 snake plot showing extracellular loops and transmembrane domains (upper portion); important residues are highlighted. Glu160, Arg189 and His269 were mutated in this study. f, Sequence alignment of GPR4, GPR65 and GPR68 to CXCR4 (PDB code 3ODU) (PROMALS-3D) was manually refined to reduce gaps and to position conserved residues. TM, transmembrane regions; IL, intracellular loop; EL, extracellular loop. Conserved residues highlighted in blue by degree of conservation while red boxes indicate residues important for receptor function. Red stars indicate residues mutated in this study. g, Sampling different regions for lorazepam binding modes in GPR68. Yellow and grey surfaces contour the binding site of 1T1t and CVX15 in CXCR4 crystal structures (PDB codes 3ODU and 3OE0, respectively), while green and red surfaces sample the entire binding pocket. The magenta surface represents the canonical orthosteric biogenic amine site. h, ZINC32547799 in its predicted orientation and interactions with GPR68. i, Optimization of ogerin (magenta, thin lines) to C2 (brown, structure in Fig. 3a) by insertion of a single methylene is predicted to improve packing in the aryl pocket of the ogerin site. Adding a second methylene, thus creating a propyl linker in C3 (yellow, structure in Fig. 3a), is predicted to disrupt the packing and thus to reduce the allosteric effect.
Extended Data Figure 3 Heat map of off-target activities of lead compounds at potential CNS drug targets.
Radioligand binding assays were carried out by the National Institute of Mental Health Psychoactive Drug Screening Program (NIMH PDSP) as described previously56,57 (online protocols available at http://pdsp.med.unc.edu/pdspw/binding.php). Values represent mean binding affinities (pKi, n = 2–4). Affinities lower than a pKi of 5, or less than 50% inhibition at 10 μM, are shown as a minimum of 5 on the pKi scale. The hERG inhibition activity was tested in a hERG functional assay as previously published59. AMPA, aminomethylphosphonic acid receptor; BZP, benzodiazepine receptor; DAT, dopamine transporter; DOR, delta (δ) opioid receptor; KA, kainate acid receptor; KOR, kappa (κ) opioid receptor; MOR, mu (μ) opioid receptor; NAT, noradrenaline transporter; NMDA, N-methyl-D-aspartate receptor; ND, not determined; PBR, peripheral benzodiazepine binding site; SERT, serotonin transporter; hERG, human ether-a-go-go-related gene (potassium channel Kv11.1).
Extended Data Figure 4 Confirmation of modelling results via mutagenesis.
a, b, Protons showed agonist activity at GPR68 wild-type and mutant receptors in cAMP production (a) and calcium release (b); parameters are in Supplementary Table 4. c, Relative GPR68 wild-type and mutant receptor expression levels determined by anti-Flag immunoblotting (n = 3). d, Proton-mediated cAMP production in untransfected cells (n = 16). e, Calcium release by lorazepam and selected ZINC compounds (10 μM at pH 8.0, n = 6–22 measurements). f–j, Effect of ogerin and ZINC32547799 (10 μM) on proton-mediated cAMP production (f and g, n = 4), calcium release (h and i, n = 3), and phosphatidylinositol hydrolysis (j, n = 3) at GPR68 wild-type or mutant-transfected HEK293T cells. k, Effect of ogerin and ZINC32547799 on phosphatidylinositol hydrolysis at pH 8.4 at GPR68-transfected GPR68 HEK293T cells (n = 3). Normalized results represent mean ± s.e.m. and curves were analysed using a four-parameter logistic function.
Extended Data Figure 5 Control experiments for signalling and pharmacology.
a, Basal cAMP production of GPR68 wild-type and mutant receptors (mean ± s.e.m., n = 24–46 measurements). b, pH-dependent activity of ogerin at GPR68 wild type (mean ± s.e.m., n = 3). c, Ogerin concentration–responses at GPR68 wild-type and mutant receptors at pH 9.0 (c, mean ± s.e.m., n = 3), under which cAMP reporter assay was not affected (d–f). d–g, Proton modulated isoproterenol-mediated Gs-activation via β2-adrenergic receptors in untransfected (d, f) and GPR68-transfected (e, g) cells. Normalized results (basal at pH 9.5 for d and e; or corresponding buffer control for f and g) represent mean ± s.e.m. (n = 6). h, i, Inverse agonist and antagonist activity (Ki of 220 nM) of ogerin at A2A (cAMP production, h) and weak antagonist activity (Ki of 736 nM) at 5-HT2B receptors (calcium mobilization, i). 5′-N-ethylcarboxamidoadenosine (NECA) and 2-chloro-N6-cyclopentyladenosine (CCPA) served as agonist controls, while CGS15943 is an inverse agonist control for A2A receptors. Normalized results represent mean ± s.e.m. (n = 3). Curves were analysed in GraphPad Prism with the built-in four-parameter logistic function. j, k, Lead compounds (10 μM) showed no agonist (j) or antagonist (k) activity at CXCR4 receptors (cAMP production) with CXCL12 as an agonist control (1 or 3 μM) or AMD 3100 (10 μM) as an antagonist control. Results represent mean ± s.d. (n = 2).
Extended Data Figure 6 Primary screening and comparison of allosteric parameters of 13 ogerin analogues at GPR68.
The 13 ogerin analogues (structures in Supplementary Table 9) identified from docking a virtual library of more than 600 ogerin derivatives were synthesized (Supplementary Information). a–e, Production of cAMP was measured in transiently transfected HEK293T cells at 10 μM and five different pH conditions, pH 8.4 (a); pH 7.9 (b); pH 7.4 (c); pH 7.0 (d); and pH 6.5 (e), to reveal any pH-dependent potentiation activity. Normalized results represent mean ± s.e.m. (n = 8–16 measurements). f, Graphic comparison of the allosteric parameters logα and logβ. Proton concentration–responses were carried out in the absence and presence of increasing concentrations of ogerin and its analogues, results were analysed using a standard allosteric operational model to obtain allosteric parameters. Values represent mean ± s.e.m. (n ≥ 3; see details in Supplementary Table 8).
Extended Data Figure 7 Characterization of potent GPR68 PAMs.
a–c, Concentration–response curves of H+ in the absence and presence of increasing concentrations of CGH2466 (a, a′), tracazolate (b, b′) and SLV320 (c, c′) and in the absence (left column, a, b, c) and presence (right column, a′, b′, c′) of phosphodiesterase inhibitor (Ro 20-1724, 30 μM) at GPR68-expressing cells. Normalized results (mean ± s.e.m., n = 8 for CGH2466; n = 5 for tracazolate; n = 5 for SLV320 for left column and n = 3 for right column) were analysed using a four-parameter logistic function and the standard allosteric operational model (not shown). Allosteric parameters in absence of Ro 20-1724 are summarized in Supplementary Table 8. For each pair of fittings, the proton potency value (negative logarithm of the half-maximum effective concentration (pEC50)) from the agonist concentration–response curve (right) in the absence of testing compound was used as the pKA for the allosteric operational model (left). d, Schematic showing the shared pharmacology among GABAA, adenosine GPCRs and GPR68 ligands. Molecules along each edge of the triangle have been shown to have activity at both targets, whereas tracazolate, in the middle, shows activity at all three.
Extended Data Figure 8 GPR68 mouse biology.
a, Ogerin (Og) and lorazepam (Lo) activate PKA and p42/p44 MAP kinase in HEK293 cells stably expressing haemagglutinin (HA)-tagged GPR68 but not HApcDNA; vehicle (Ve). b, c, GPR68 knockout (n = 7) mice exhibited no differences in contextual memory retrieval (b) or cued memory retrieval (c) as compared to wild-type mice (n = 8). d–f, At 10 mg kg−1, the ogerin isomer ZINC32547799 had no effect on learning (d) or contextual and cue memory (e, f) in wild-type mice (vehicle, n = 6; drug, n = 7). g–i, At 30 mg kg−1, ZINC32547799 enhanced wild-type learning (g, drug × time interaction, F(3.39) = 3.58, P = 0.022; drug alone F(1.39) = 1.19, P = 0.295; Bonferroni post-hoc test revealed a significant effect (P < 0.05) at the third unconditioned/conditioned stimulus training point, two-way ANOVA), but had no effect at contextual and cue memory (h, i) (vehicle, n = 7; drug n = 8). Male mice at age of 6–8 weeks were used in the test. Normalized contextual memory retrieval (d) and cued memory retrieval (f) are presented in Fig. 4c, d.
Extended Data Figure 9 Screening of ZINC compounds predicted to be active at GPR65 based on BTB09089 docking poses.
a–d, Primary screening with ZINC compounds (30 μM) for agonist activity at GPR65 when receptors were kept inactive at pH 8.40 (a); at control HEK293T cells for nonspecific activity (b); at GPR65 when receptors were activated at pH7.40 for modulator or antagonist activity (c); at GPR65 when receptors were activated by BTB09089 (30 μM) at pH 8.40 for modulator or antagonist activity (d). Normalized results represent mean ± s.e.m. from a minimum of three assays (each in minimum of triplicate and a total of ≥16 measurements). The red dashed line in b–d indicates the 20% inhibition line (an arbitrary cut-off line).
Extended Data Figure 10 Characterization of GPR65 allosteric modulators at wild-type and mutant receptors.
a, BTB09089 showed weak agonist activity, but failed to potentiate proton activity at GPR65 (n = 8). b, Selected compounds from Extended Data Fig. 9b, c were tested for GPR65 specific inhibition (n = 16–56 measurements). Several compounds (such as ZINC41613384, ZINC9468042 and ZINC62678696) showed GPR65-specific inhibition. c, Selected compounds from Extended Data Fig. 9b, d were tested for antagonist activity against BTB09089-activated signal at GPR65 (n = 16–64 measurements). ZINC62678696 showed GPR65 specific inhibition when it was activated by either proton or BTB09089. d, ZINC62678696 inhibited GPR65 activity. e–g, Proton concentration–responses (e), BTB09089 concentration–responses (f), and ZINC13684400 concentration–responses (g) at GPR65 mutant receptors. Normalized results represent mean ± s.e.m. (n ≥ 3) and curves were analysed in GraphPad Prism with a standard four-parameter logistic function. Corresponding curves of proton at GPR65 wild-type receptors (from Extended Data Fig. 4a) and BTB09089 and ZINC13684400 (from Fig. 5e) are also included (dashed lines) for comparison. Pharmacological parameters are listed in Supplementary Table 13.
Extended Data Figure 11 In vivo behavioural profiling of GPR68-knockout mice.
a, b, No effects of GPR68 deletion on distance travelled in an open field. Data represent mean ± s.e.m. for each group for a one-hour test session. c, d, No difference on latency to fall from an accelerating rotarod. Data represent mean ± s.e.m. for each group. e–h, Decreased startle responses in GPR68 knockout mice after presentation of acoustic stimuli (e, f). Data represent mean ± s.e.m. for each group. No effects of genotype were found for levels of prepulse inhibition (g, h). Data represent mean ± s.e.m. for each group (*P < 0.05). i–l, No difference at acquisition and reversal learning in the Morris water maze. Data represent mean ± s.e.m. of four trials per day. Subject numbers were 9 wild-type and 7 knockout male mice, and 12 wild-type and 11 knockout female mice.
Supplementary information
Supplementary Methods
This file contains Supplementary Methods ‘Chemistry procedures for ogerin and its analogues’ and additional references. (PDF 537 kb)
Supplementary Tables
This file contains Supplementary Tables 1-13 and additional references. (PDF 2596 kb)
Supplementary Data
This zipped file contains 4 PDB files as follows: for modeled complex of GPR65 with BTB09089; for modeled complex of GPR65 with both BTB09089 and ZINC62678696; for modeled complex of GPR68 with lorazepam; and for modeled complex of GPR68 with ogerin. (ZIP 230 kb)
Supplementary Figure 1
This file contains Supplementary Figure 1 ‘Yeast-based screening to identify potential ligands for GPR68’, which is a heat map showing growth responses of yeast expressing 24 different GPCRs (horizontal axis) to 446 compounds in the NCC library (vertical axis). Blue color is 0% (or less than 0%) above background; white is 100% above background, red color indicates 200% (or more) above background growth stimulation, i.e., agonist activity. Data were clustered using GENE-E software (http://www.broadinstitute.org/cancer/software/GENE-E/) using complete linkage algorithm. GPR68 is the right-most column in the heat map and details are shown in Figure 1a. (ZIP 172 kb)
Rights and permissions
About this article
Cite this article
Huang, XP., Karpiak, J., Kroeze, W. et al. Allosteric ligands for the pharmacologically dark receptors GPR68 and GPR65. Nature 527, 477–483 (2015). https://doi.org/10.1038/nature15699
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature15699